what portion of a trna molecule binds to the codon of an mrna molecule?
What is tRNA?
Definition of tRNA- Transfer RNA or tRNA is a type of RNA molecule that helps to decode information present in mRNA sequences into specific proteins.
- tRNA molecule is a carrier of amino acid that brings appropriate amino acrid to ribosome based on the codon present in mRNA sequence.
- tRNA is also known as an adaptor molecule every bit it translates the codons present in mRNA sequences into amino acids.
- tRNA is simply 70-90 nucleotides in length, making information technology the smallest out of the iii primary RNAs (mRNA and rRNA).
- The molecular weight of the tRNA molecule ranges from 25,000 to thirty,000 Dalton.
- tRNA is encoded by DNA in the jail cell nucleus and transcribed with the assist of RNA polymerase ΙΙΙ.
Backdrop of tRNA
- tRNA can make hydrogen bonds with mRNA and besides can grade ester linkage with amino acids, thus linking mRNA with the amino acid at the site of poly peptide synthesis.
- tRNA bond complementary with each iii-nucleotide of mRNA chosen codons and this happens in an antiparallel style. The function of tRNA that binds with the codon of mRNA is termed as anticodon arm of tRNA.
- This complementary base pairing of tRNA and mRNA makes the correct translation of the correct amino acid that is inserted into the growing polypeptide chain.
- The tRNA is a single-stranded RNA molecule that has the proximity to fold upon itself and creates an intra complementary base of operations pairing which gives enhance to hydrogen-bonded stems and associated loops that contains nucleotides with modified bases.
- In addition to the usual bases i.e., Adenine, Uracil, Cytosine, and Guanosine, tRNA also contains modified unusual bases like for Adenine, in that location is an unusual base of operations chosen Inosine, too in place of Uracil, there are Psuedouracil and Dihydrouridine.
- These modified unusual bases and intramolecular base pairing of tRNA brand it stable and protect it from getting degraded by the enzyme RNase.
Kinds of tRNA Structure
The tRNA has three kinds of structure:
ane. Chief structure
- A linear single strand of ribonucleotides running in a 5′ to 3′ management.
- The number of ribonucleotides ranges from sixty-90, the most normally found length is 76 ribonucleotides.
- A unmarried tRNA molecule having a master construction consists of about twenty% of modified bases.
- Based on these modified bases, the linear sequence of tRNA molecule can be divided into three arms: D arm containing a modified base Dihydrouridine, Anticodon arm containing unusual bases similar Inosine derived from Adenine, or Pseudouridine from Uracil or Lysidine from Cytosine, TΨC arm containing modified base Pseudouridine.
- In add-on to these three artillery with modified bases, an arm is present at the three′ terminal usually containing CCA nucleotide and is referred to equally Acceptor arm that acts as an zipper site for incoming amino acids.
- D arm serves every bit a recognition site for the enzyme aminoacyl tRNA synthetase.
- The Anticodon arm binds with the codon of mRNA during protein synthesis.
- TΨC arms comprise a ribosome binding site.
2. Secondary structure (Cloverleaf model)
- The secondary construction of tRNA is of 74-95 nucleotides sequences that are stable and fold on itself to give a clover leaf-like structure containing four arms sometimes fifty-fifty v in longer tRNAs.
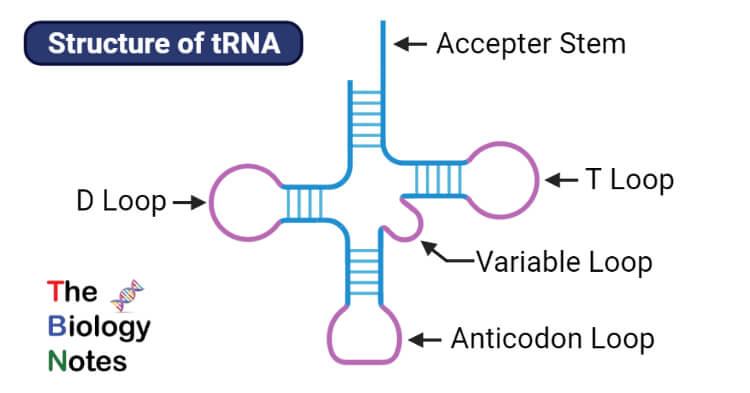
- These iv base of operations-paired artillery are namely, Acceptor arm, D-arm, TΨC arm, and Anticodon arm.
- The acceptor's arm is a 7 to ix base pair in length, this arm is formed by base of operations pairing of 5′ terminal and iii′ concluding nucleotides.
- Out of these four base of operations-paired arms, 3 arms finish up in a loop due to unpaired base-pairing forming D- loop, Anticodon loop, and TΨC loop respectively. However, the acceptor arm does not class a loop but contains an extension of the free 3′ hydroxyl group comprising of CCA nucleotides that are used to attach amino acids.
- The carboxylic group of amino acids joins with costless iii′ hydroxyl of Adenine in CCA to aminoacyl tRNA, the procedure is also referred to as charging of tRNA and is catalyzed by an enzyme, aminoacyl tRNA synthetase.
- The acceptor arm is located reverse to Anticodon arm.
- D arm consists of a stem that is near four to five base pairs in length which finish in a loop that usually contains modified pyrimidine base nucleotides, Dihydrouridine.
- D arm serves equally a recognition site for the enzyme aminoacyl tRNA synthetase.
- Anticodon arm consists of a stem that is about 6 base pairs in length which end in a loop due to vii unpaired bases. Out of which three bases class an anticodon.
- This first anticodon base is sometimes modified with unusual bases similar Inosine derived from Adenine, or Pseudouridine from Uracil, or Lysidine from Cytosine.
- This anticodon base of operations of the anticodon arm recognizes codon in mRNA and binds to it during protein synthesis.
- The TΨC arm consists of a stem that is five base pairs long which ends in a loop that further contains 7 unpaired base pairs, usually containing Thymine, Pseudouridine and Cytosine hence the name TΨC arm.
- The TΨC loop contains a ribosome bounden site.
- In addition to four base-paired arms, a variable arm can usually be nowadays in long tRNAs. These variable arms vary in length from three-21 nucleotides depending on the type of amino acid they code.
- The variable arm lies between the anticodon arm and TΨC arm, and is usually short and looks sort of different from the regular four artillery. The base of operations pairing is rarely seen in this arm, so information technology appears as loops due to unpaired bases.
- The variable arm accounts for the stability of a tRNA molecule.
iii. Tertiary structure
- It is a three-dimensional structure, where tRNA adopts an L-shaped structure.
- This L-shape is formed when the stem of acceptor and TΨC arms stack at i side and stem of anticodon and D artillery stack on another side and both forms extended helices.
- Both of these extended helices align at a right angle, and in doing so, the D-loop and TΨC loop align together.
- The tertiary structure is the about stable structure out of all, and this stability is acquired due to hydrogen bonding in between nitrogenous bases and likewise between nitrogenous bases and the ribose-phosphate courage.
- To synthesize proteins by reading codon in mRNA a tRNA molecule must have a 3rd structure.
Composition of tRNA
- tRNA being a blazon of RNA molecule is composed of a ribose sugar, phosphate, and nitrogenous base.
- It is a biopolymer of ribonucleotides.
- tRNAs are transcribed from the genes present in DNA. And every bit we know, Dna runs from a v′ end to a 3′ terminate, and then tRNA that is synthesized does have the same direction.
- During the processing of the forerunner tRNA to mature tRNA, many additions and modifications are made, hence
- These modified bases in a tRNA construction characterize distinct portions in the chains of ribonucleotides.
- Based on these modified bases, tRNA molecule has three arms or telephone call it segments, one being a D arm that distinctively has an unusual base chosen Dihydrouridine derived from the methylation of Uracil, some other ane beingness TΨC arm that is characterized by the presence of some other methylated Uracil called Pseudouridine and the concluding 1 is Anticodon arm that has a modified base of Adenine, called Inosine.
- Along with these 3 distinct arms, an important arm is also nowadays that does not contain modified bases but after processing becomes an acceptor arm that attaches amino acids.
- In summary, a tRNA molecule is composed of bondage of ribonucleotides, in which certain modification of bases and improver of nucleotides takes identify during processing resulting in the germination of four arms: D arm, TΨC arm, Anticodon arm, and Acceptor arm.
Processing of tRNA
- Like all RNA, tRNA is synthesized by transcribing the genes encoded in the DNA molecule with the help of an enzyme called RNA polymerase ΙΙΙ.
- And are fabricated translation-ready (matured) through splicing, CCA improver, and methylation of bases in nucleotides.
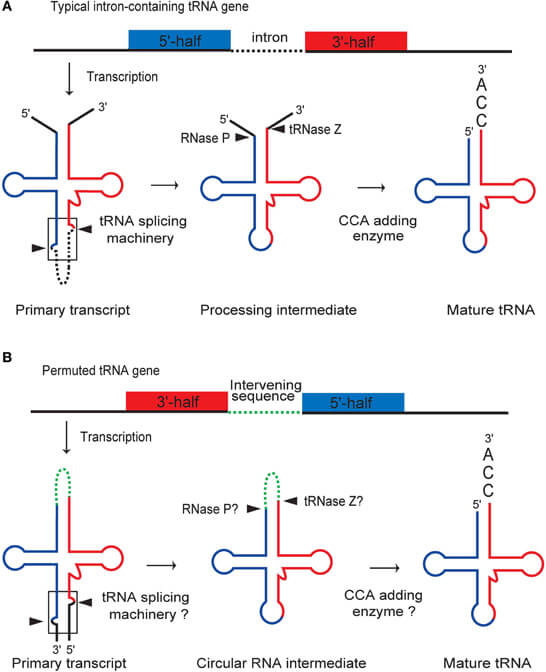
- The newly transcribed precursor tRNA has extra sequences at the v′ and iii′ ends that are broken off past the action of an enzyme chosen RNase P, which is a ribonuclease.
- In some cases, precursor tRNA likewise contains a not-coding region called introns, which are spliced out past a specific endonuclease, and and so the remaining fragments are joined with aid of RNA ligase.
- CCA addition at the 3′ end is done with the help of an enzyme called tRNA nucleotidyltransferase.
- Modification of bases besides takes place during the processing of precursor tRNA and is made possible past an enzyme, tRNA methylase that helps to provide a methyl donor, thus making the bases methylated which gives rise to modified unusual bases like Pseudouridine, Inosine, Dihydrouridine, and Lysidine.
- Each modification is fabricated to functionalize tRNA for protein synthesis.
- Addition of CCA nucleotides at three′ end aid to attach specific amino acid by forming ester bail (esterification) with a free hydroxyl group of Adenine nucleotide of CCA. This reaction is catalyzed by an enzyme aminoacyl tRNA synthetase.
- Splicing makes the tRNA free from introns. And methylation of bases makes them recognizable by aminoacyl tRNA synthetase and ribosome.
- Hence, all this processing must take place, to make mature tRNA that can further be benign during protein synthesis.
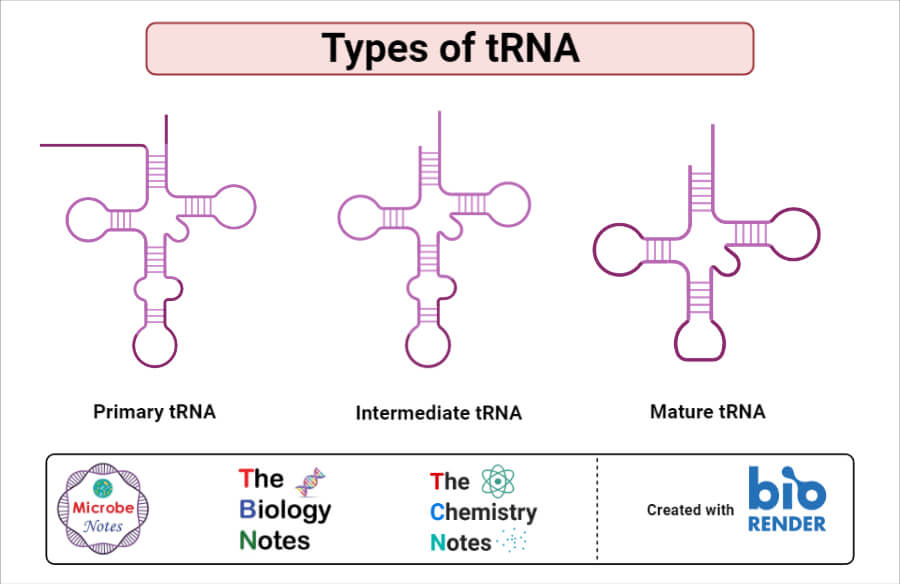
Types of tRNA
- The types of tRNA are categorized on the types of amino acids it carries. A human being typically has twenty essential amino acids which thus requite rise to 20 different types of tRNA.
- However, alternatively, they can besides be categorized based on their anticodons.
- When genes are expressed to proteins. DNA is transcribed to mRNA which is then translated into poly peptide by tRNA. Deoxyribonucleic acid contains four nucleotides so their possible combination gives ascent to 64 codons.
- Out of 64 codons, 3 codons are finish codons i.due east., UAG, UAA, and UGA. These three codons trigger the termination of translation. In addition to tRNA molecules beingness needed to pair with each amino acid. Some other tRNA molecules are also needed to pair upward with these terminate codons to halt the protein synthesis.
- Some codons lawmaking for the aforementioned amino acid, thus the real number of tRNA is not 64. This redundancy in genetic code is referred to as "wobble".
- Due to this back-up, very few species of organisms take exactly 61 codons that code for amino acids.
- tRNA interacts with codons via its anticodon loop. Base pairing between anticodon and codon ensures the specificity of amino acids to be incorporated in polypeptide chains.
Functions of tRNA
- tRNAs are vital molecules taking role in protein synthesis that transport amino acids to the ribosome where these amino acids are added according to the codon sequences encoded in the mRNA molecule.
- Picks upwards specific amino acids in the cytoplasm and carries them to the site of poly peptide synthesis.
- Also participate in not-protein constructed processes, such as primer during the reverse transcription of retroviruses.
- Information technology is also known equally an adaptor molecule because it attaches specific amino acids that the codon represents in the mRNA sequence. It tin also be referred to as a bridge that links amino acids and nucleotides.
- Mutations in tRNA or tRNA processing are linked with various human diseases, similar cancer and neurodegenerative diseases.
- In recent times, the tRNA molecules are contradistinct to accommodate desired anticodons that result in translating unusual amino acids that do some specific functions. For case, the tRNA-dependent aminoacylation of bacterial jail cell membrane lipids has deemed for increased virulence of bacteria and their resistance against antibiotics.
tRNA Frequently Asked Questions (FAQs)
Q. What is tRNA?
Ans. Transfer RNA or tRNA is a type of RNA molecule that helps to decode information present in mRNA sequences into specific proteins.
Q. What is the limerick of tRNA?
Ans. tRNA is composed of a ribose sugar, phosphate, and nitrogenous base of operations.
Q. What are the 3 kinds of tRNA structure?
Ans. tRNA can be divided into iii kinds of structures- Primary construction, Secondary structure (Cloverleaf model), and Third structure.
Q. How many tRNAs are there in humans?
Ans. There are 22 unlike tRNAs in homo mitochondria.
Q. Where is the tRNA produced?
Ans. In Eukaryotes, Mature tRNA is produced (generated) in the Nucleus.
Q. What are the functions of tRNA?
Ans. tRNA helps in protein synthesis that transports amino acids to the ribosome where these amino acids are added according to the codon sequences encoded in the mRNA molecule.
References
- Soma A. Circularly permuted tRNA genes: their expression and implications for their physiological relevance and development. Forepart Genet. 2014 Apr 1;5:63. doi: 10.3389/fgene.2014.00063. PMID: 24744771; PMCID: PMC3978253.
- Hori, H., Tomikawa, C., Hirata, A., Toh, Y., Tomita, K., Ueda, T. and Watanabe, K. (2014). Transfer RNA Synthesis and Regulation. In eLS, John Wiley & Sons, Ltd (Ed.).
- https://www.britannica.com/science/transfer-RNA
- https://biologydictionary.internet/trna/
- https://www.nature.com/scitable/definition/trna-transfer-rna-256/
- https://courses.lumenlearning.com/microbiology/chapter/construction-and-function-of-rna/
- https://www.slideshare.internet/manjeshsaakre/trna-construction-and-role
- http://dnaofbioscience.blogspot.com/2016/05/clover-leaf-model-of-t-rna.html
- https://en.wikibooks.org/wiki/Structural_Biochemistry/Nucleic_Acid/RNA/Transfer_RNA_(tRNA)
- https://bio.libretexts.org/Bookshelves/Microbiology/Book%3A_Microbiology_(Boundless)/7%3A_Microbial_Genetics/7.06%3A_Translation%3A_Protein_Synthesis/seven.6A%3A_Processing_of_tRNAs_and_rRNAs
- https://courses.lumenlearning.com/wm-biology1/chapter/reading-trna/
- O'Donoghue, P., Ling, J., & Söll, D. (2018). Transfer RNA function and evolution. RNA biology, fifteen(4-v), 423–426. https://doi.org/10.1080/15476286.2018.1478942
- Kirchner, S., Ignatova, Z. Emerging roles of tRNA in adaptive translation, signaling dynamics, and disease. Nat Rev Genet 16, 98–112 (2015). https://doi.org/10.1038/nrg3861
Source: https://thebiologynotes.com/transfer-rna/
0 Response to "what portion of a trna molecule binds to the codon of an mrna molecule?"
Enregistrer un commentaire